Protein Interactions
Introduction
Protein interactions inform how biological processes and structures function within cells. Protein interaction networks - like those generated by STRING - can summarize known connections between clusters of proteins, and can also help predict novel interactions [1]. The mapping of these relationships serves as a scientific foundation for deepening the understanding of the proteome [2].
Results
Discussion
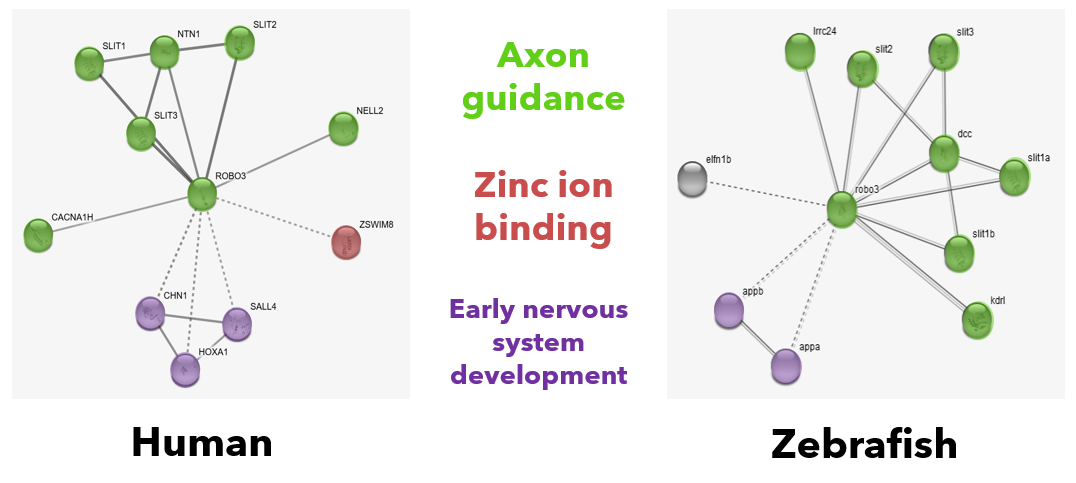
STRING generated an output for the primary relationships between ROBO3 and other proteins in both humans and Danio rerio zebrafish. The relationship clusters of GO terms (represented by color code) are very similar, further supporting the homologous function of ROBO3 within my model organism species. These identified gene clusters are unsurprising given the known role of ROBO3 within the biological process of axon guidance. This process functions within the neurons of the developing nervous system, and communication between neurons often involves zinc ions - which explains the zinc ion binding GO term. These GO clusters work closely together to contribute to the phenotypes shown in synesthesia and other neurological disorders associated with ROBO3 mutations.
References
- STRING. http://string-db.org/
- Adams, J. The Proteome: Discovering the Structure and Function of Proteins. Nature Education, (3):6 (2008).